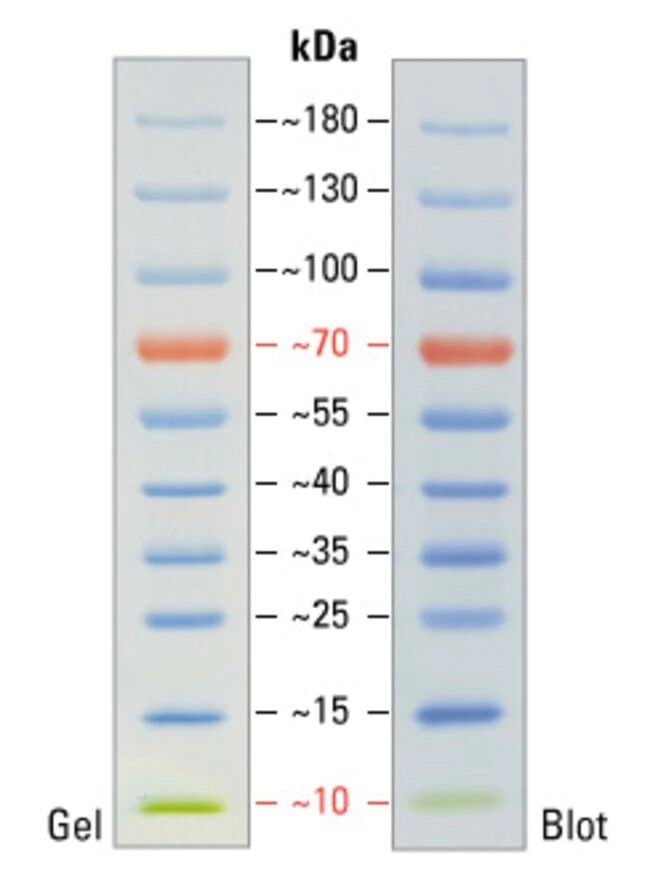
If the target protein is of low molecular weight a high percentage gel should be chosen in order to maximize protein resolution. Selecting the correct gel for the target protein can be a crucial step in optimizing detection and quantification. In general Bio-Rad recommends its Criterion™ Tris-tricine precast gels for small proteins and its Criterion XT Tris-acetate precast gels for separating higher molecular weight proteins (see section below for more details). Knowing the size of the target protein will indicate the gel that should be chosen and the conditions for transfer. One disadvantage of protein precipitation is that proteins might denature, making the pellet difficult to re-solubilize. After centrifugation of the precipitated protein to a pellet, the supernatant is removed and the protein pellet is re-dissolved in sample buffer. In addition to a nuclear protein extraction Bio-Rad has protein extraction kits for membrane bound proteins as well as cytosolic signaling proteins.Īnother strategy for increasing specific protein detection is to add a compound such as acetone that causes protein to precipitate. An anti-lamin A/C antibody ( VMA00200) can be used as a marker to differentiate nuclear from cytoplasmic samples. For recovery of nuclear proteins, Bio-Rad offers the ReadyPrep™ Protein Extraction Kit (Cytoplasmic/Nuclear). If the target protein is primarily expressed in a single cellular compartment (membrane or nucleus) then recovery of the proteins from the individual component will increase the chances of quantifiable detection. Recovery of protein from individual cellular compartments Reduce Cu 2+ to Cu + which binds to protein Under acidic conditions Coomassie Brilliant Blue G-250 binding to protein Peptide bonds react with copper under alkaline conditions to produce Cu + They are all colorimetric assays and can utilize known concentrations of Bovine Serum Albumin (BSA) to construct a standard curve against which samples can be evaluated. There are three primary methods for determining the concentration of recovered protein. Approximate Protein Recovery (based on HeLa cells, the number of cells will vary according to cell type).ĭetermining concentration of recovered protein 
Typical loading amounts of protein added to each well range from 10 µg (~330,000 cells) to 50 µg (~1,650,000 cells). HeLa cells contain approximately 300 pg per cell. Protein recovery can vary according to the cell type and recovery method. However, please note that using an insufficient amount of lysis buffer may lead to an incomplete lysis of the sample. This will allow the addition of increased amounts of protein per well furthering chances of detecting the low abundant protein. When lysing the cell or tissue, use as little lysis buffer as possible in order to concentrate samples as much as possible. 
Use of the correct lysis buffer will improve the detection of the low abundant target protein. Non-ionic detergents induce less protein denaturation so if their use is often recommended with primary antibodies raised against native proteins. If further denaturing is required a reducing agent (DTT or Beta mercaptoethanol) may be used to disrupt S-S bonds. As such their use with antibodies raised against denatured forms or synthetic peptides of the target protein is often recommended. Ionic detergents are considered harsher than the non-ionic detergents because of their ability to interfere with protein structure. The best known of the ionic detergents is sodium dodecyl sulfate (SDS) while non-ionic detergents include Tween, Triton X and NP-40. The primary active agent in lysis buffer is the detergent which can be ionic or non-ionic. Lysis is the process in which the cell membrane is broken and the contents of the cell are re-suspended in a soluble form. There are several steps for maximising the recovery of your target protein from the sample. Initial improvements in the detection and quantification of low abundant proteins can be made at the sample preparation stage. As such detection of proteins with low expression levels is a common problem for researchers. To help you optimize both the detection and quantification of such proteins and overall quality of the final western blot image, we have put together useful tips for the following steps: The technique of western blotting illuminates molecular events including protein expression, protein localisation, protein-protein interactions or post translational modifications (PTMs). These molecular events are often very subtle. For example within an individual signaling pathway only a certain percentage of the molecules may have undergone a PTM. Enhance detection and quantification of low abundance protein
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
